1990 : The Experiment That Did the Opposite of What It Should Have
In 1990, Carolyn Napoli, Christine Lemieux, and Richard Jorgensen at DNA Plant Technology
Corporation set out to do something straightforward: make purple petunias more purple.
Building the construct
To overexpress chalcone synthase (CHS) the key enzyme in anthocyanin pigment biosynthesis
they needed a promoter that would push transcription as hard as possible. They chose the CaMV
35S promoter, derived from Cauliflower mosaic virus. This promoter is constitutively active in
virtually all plant tissues and drives transcription at very high levels because it was originally
evolved by the virus to hijack the plant's own transcriptional machinery for viral replication. It
contains a tandem repeat enhancer element that recruits host RNA polymerase II with unusually
high efficiency. In molecular biology, it became the standard tool for maximal gene expression in
plant systems.
The construct was assembled by fusing this 35S promoter to the coding sequence of a petunia CHS
cDNA clone, with a nopaline synthase (nos) polyadenylation signal at the 3' end to ensure proper
transcript termination and stability. When RNA polymerase finishes transcribing a gene, the
transcript needs a defined endpoint. The nos terminator provides the polyadenylation signal
(AAUAAA or a variant), which tells the cell's cleavage/polyadenylation machinery to cut the pre-
mRNA at a specific site downstream and add a poly(A) tail — a stretch of ~200 adenine
nucleotides. This poly(A) tail does two things: it protects the 3' end of the mRNA from
exonucleases that would otherwise chew it up, and it recruits poly(A)-binding proteins (PABPs) that
cooperate with the 5' cap to promote ribosome recruitment and efficient translation. Without a
functional terminator, transcription can run through into downstream sequences, producing aberrant
read-through transcripts that are unstable, poorly translated, or that interfere with neighboring
genes. The nos terminator was borrowed from Agrobacterium — it originally terminates the
nopaline synthase gene in the T-DNA — and it works reliably in plant cells because the core
polyadenylation machinery is conserved across eukaryotes.
__The delivery system: how do you get a gene into a plant?__
The tool is a soil bacterium called Agrobacterium tumefaciens, which already knows how to insert
DNA into plant cells — it's been doing it for millions of years.
How it works in nature. When a plant root gets wounded, the damaged cells leak acetosyringone, a
chemical signal. Nearby Agrobacterium detect it, and this switches on their vir (virulence) genes.
The Vir proteins cut out a specific stretch of DNA from the bacterium's Ti plasmid (Tumor
Inducing) — the transfer DNA « T-DNA », flanked by two 25-bp border repeats — and thread it as
a single strand into the plant cell's nucleus, where it inserts more or less randomly into the
chromosomes. In the wild, this T-DNA carries genes that do two things for the bacterium: force the
plant to grow a tumor (by producing the hormones auxin and cytokinin) and manufacture opines,
nutrients only the bacterium can eat. It's a parasitic trick — reprogram the host to feed you.
How scientists adapted it. The tumor and opine genes are stripped out ("disarmed"), and the gene
you actually want is spliced in between the border sequences. The vir machinery doesn't care what's
between the borders — it transfers whatever is there. This is the basis of almost all plant genetic
engineering.
__The vector: pJJ3942__
The CHS overexpression cassette was loaded into a plasmid called pJJ3942, built on the broad-host-
range replicon pRK290 — a backbone that can replicate in multiple bacterial species, which is
essential here because the vector needs to be maintained first in E. coli (for cloning) and then in
Agrobacterium (for plant transformation). This is a binary vector, meaning it's one half of a two-
plasmid system:
• The binary vector (pJJ3942) carries the gene of interest between the T-DNA borders. This
is the part that gets transferred into the plant.
• The helper plasmid, already resident in the Agrobacterium host, supplies the vir genes —
the transfer machinery.
Why split them? So you can freely modify the gene-carrying plasmid without breaking the transfer
system. Swap genes, add markers, redesign the cassette — the vir functions stay untouched on the
other plasmid.
Besides the CHS cassette, pJJ3942 carried two additional elements: a CaMV 35S enhancer near the
right T-DNA border, and an NPTII gene (neomycin phosphotransferase II) driven by a nos
promoter, which confers kanamycin resistance — the selectable marker used to identify transformed
cells later.
__Getting the construct into Agrobacterium__
The vector was introduced into the disarmed A. tumefaciens strain LBA4404 through triparental
mating: a conjugation procedure where a helper E. coli strain acts as a go-between, shuttling the
binary vector from the cloning host into Agrobacterium.
Transforming the petunias
Leaf pieces from three petunia genotypes — Pink Cascade, R18, and V26 — were wounded and
incubated with the engineered bacteria in the presence of acetosyringone (to activate the vir genes).
After two days of co-cultivation, the explants were moved to selection medium.
__Selection: how do you know which cells took up the gene?__
The medium contained kanamycin at 300 mg/L — a concentration that kills untransformed plant
cells. Here's why it works:
Kanamycin is an aminoglycoside antibiotic that blocks protein synthesis by binding the 30S
ribosomal subunit. Plant chloroplasts and mitochondria use bacterial-type 70S ribosomes, making
them direct targets. In untransformed cells, kanamycin shuts down organellar translation, the
chloroplasts fail, the cells bleach, and growth stops.
Cells that successfully integrated the T-DNA, however, express the NPTII enzyme, which
phosphorylates kanamycin and renders it inactive. These cells survive, divide, and regenerate into
shoots. After rooting, the plantlets were transferred to soil and grown in a greenhouse.
__The unexpected result__
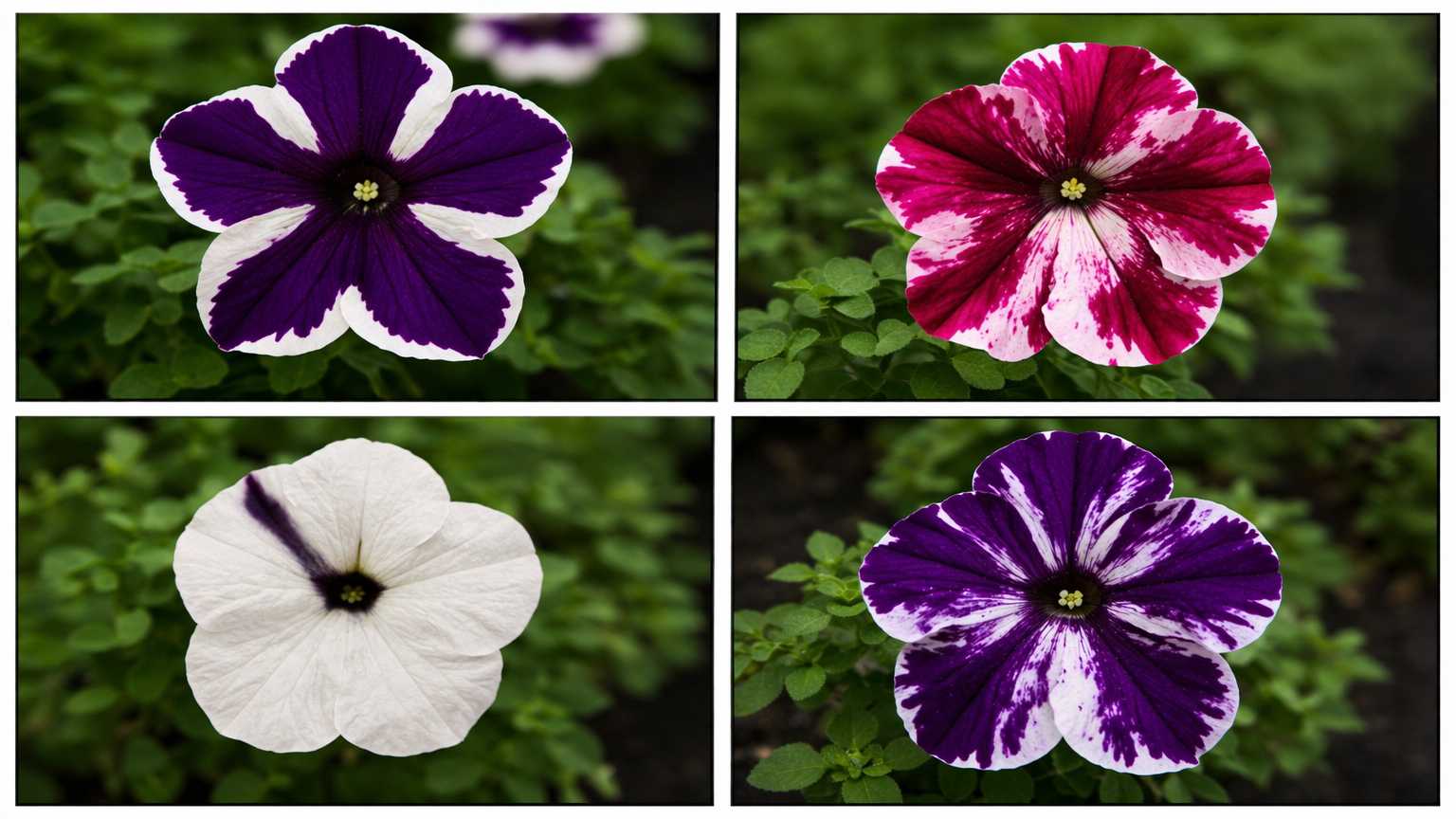
Instead of darker flowers, 42% of transgenic plants produced white or partially white flowers. Not a
single plant got darker. The result held across all three genetic backgrounds and dozens of
independent insertion events, ruling out a fluke. Among hundreds of control transgenotes carrying
other constructs, none showed altered flower color.
The white sectors weren't clonal they didn't follow cell lineage boundaries which meant this wasn't
a simple mutation. Something dynamic was happening at the cellular level. Two distinct pattern
classes emerged: a "wedge" pattern and a radial "Cossack dancer" pattern, with variants ranging
from highly regular to chaotic "tie-dyed" arrangements.
__What the molecular data showed__
Using RNase protection assays, the team could distinguish mRNA from the transgene and the
endogenous CHS gene. In white flowers, endogenous CHS mRNA was reduced about 50-fold yet
the gene was still turned on and off at the right developmental times. The silencing wasn't shutting
down the gene's regulation, it was destroying the mRNA after it was made.
The clearest evidence came from plant 218.41, which produced solid white flowers for four months
before spontaneously growing a branch with violet flowers. This reversion was discontinuous it
appeared suddenly on a side branch immediately after branching, rather than gradually fading in.
Cuttings from each branch stably maintained their respective colors for over six months.
The seasonal context adds an important detail: the authors noted that white sectors were typically
larger in summer and smaller in winter across many transgenotes.
This seasonal variation together with the observation by Van der Krol and colleagues that
supplementary light in Amsterdam could induce visible white sectors in plants that appeared fully
pigmented under normal Dutch light conditions strongly suggests that
light intensity or photoperiod modulates the co-suppression effect. From what we now know about
RNAi, this makes mechanistic sense: higher light drives higher CHS transcription via UV and light-
responsive elements in the CHS promoter, which would increase mRNA flux and potentially exceed
a threshold that triggers or reinforces dsRNA production and PTGS cycling. Temperature could also
play a role, as RNA silencing efficiency in plants is known to be temperature-sensitive generallymore efficient at higher temperatures (25–30°C) and suppressed below 15°C. The spontaneous
reversion of 218.41 may therefore reflect a stochastic fluctuation in this silencing equilibrium,
amplified by environmental shifts during the growing season in Oakland, California.
In violet revertant flowers, both transgene and endogenous CHS mRNAs were 30- to 50-fold higher
than in white flowers. Both genes were suppressed and restored together. The authors coined the
term "co-suppression" but had no mechanism to explain it.
1998 : Fire and Mello find the Trigger
Eight years later, working in the nematode C. elegans, Andrew Fire and Craig Mello resolved the
mystery. They compared the silencing potency of sense RNA, antisense RNA, and double-stranded
RNA (dsRNA).
__The methodology__
The RNAs were synthesized in vitro from plasmid templates using phage RNA polymerases (T3
and T7) to generate sense and antisense strands separately. To produce dsRNA, the two
complementary strands were annealed together, and the preparation was verified for double-
strandedness. The target gene in these experiments was unc-22, which encodes a muscle structural
protein silencing it produces a visible twitching phenotype, providing a simple and unambiguous
readout.
The RNA was delivered by microinjection directly into the gonad syncytium of adult
hermaphrodite worms. The gonad syncytium is a shared cytoplasmic compartment that feeds into
developing oocytes, meaning that injected material is distributed to many future embryonic cells.
This delivery route also allowed the researchers to observe effects in both the injected adults (via
diffusion from the gonad into somatic tissues) and in the F1 progeny that developed from the treated
oocytes. They also tested injection into the body cavity and into the gut, finding that dsRNA was
effective from multiple delivery sites suggesting that the interfering signal could cross cellular
boundaries.
__The result__
The result was decisive: either single strand alone had at best a weak, variable effect. Double-
stranded RNA was dramatically more potent, producing sequence-specific silencing in both the
injected animals and their progeny. Crucially, only a few dsRNA molecules per cell were needed
far too few for simple one-to-one hybridization with target mRNAs. Something was amplifying the
signal.
This catalytic, sub-stoichiometric behavior pointed to an enzymatic degradation machinery what
we now know as the RISC complex, where a single loaded Argonaute protein can cleave multiple
target mRNAs in succession. In C. elegans and plants, RNA-dependent RNA polymerase (RdRP)
provides an additional amplification layer by using cleaved target mRNA as a template to
synthesize secondary dsRNA, which is then processed into a new wave of siRNAs. This
amplification loop explains the extraordinary potency and persistence of the silencing effect in these
organisms.Fire and Mello received the 2006 Nobel Prize for this work, and their findings retrospectively
explained the petunia co-suppression: the transgene must have been inadvertently generating
dsRNA, triggering this same catalytic pathway.
picture of worm Caenorhabditis Elegans taken by Bob Goldstein, UNC Chapel Hill http://bio.unc.edu/people/faculty/goldstein/ available on wikipedia
https://fr.wikipedia.org/wiki/C.elegans%28homonymie%29
2005 Proof That Nature Does It Too (Koseki et al.)
The co-suppression story involved transgenic plants. But does this silencing occur naturally? The
answer came from Petunia hybrida 'Red Star' a non-transgenic cultivar with star-shaped red and
white petal sectors.
Koseki and colleagues measured mRNA of six anthocyanin pathway genes in red versus white
sectors. Only CHS-A was suppressed in white tissue the other five were expressed normally.
Nuclear run-on assays confirmed the gene was still being transcribed; the block was post-
transcriptional. And Northern blots revealed CHS-A siRNAs (~21 nt) accumulating exclusively in
white sectors the direct molecular signature of active RNAi.
The final proof: infection with Cucumber mosaic virus (CMV), which carries the 2b silencing-
suppressor protein, completely abolished the white sectors. Flowers turned uniformly red and CHS-
A mRNA recovered. Block the RNAi machinery, the phenotype disappears.

AI generated picture of petunia Red Star
2012 The Genomic Architecture Behind the Pattern (Morita/Nakayama et al.)
What generates the dsRNA in a non-transgenic plant? Morita, Nakayama, and colleagues sequenced
the CHS-A locus and found the answer: both Picotee and Star cultivars carry two intact CHS-A
copies PhCHS-A1 and PhCHS-A2 arranged in tandem (head-to-tail) with 99% coding sequence
identity. This tandem arrangement likely promotes dsRNA formation through read-through
transcription from one copy into the next.
RT-PCR confirmed that precursor mRNAs from both copies were present in red and white tissues
alike the genes are transcribed everywhere. But mature, spliced mRNA was found only in
pigmented tissue. The transcripts are made, then selectively destroyed in white sectors.
Deep sequencing of small RNAs from Picotee petals (~55 million reads) showed CHS-A siRNAs
enriched over 100-fold in white tissue compared to pigmented tissue, predominantly 21-nt, mapping
to exon 2 of both CHS-A copies the classic Dicer footprint.
Genetically, crossing Picotee with a fully pigmented cultivar showed that the bicolor trait is
recessive and that the tandem CHS-A allele is necessary but not sufficient. Some F2 plants carrying
the tandem had fully pigmented flowers. A second, still-unidentified regulatory locus must control
where siRNA production is activated. The tandem itself likely arose through unequal crossing over
during the interspecific hybridization events that created P. hybrida in the 1830s the precursors of
PhCHS-A1 and PhCHS-A2 exist independently in wild P. integrifolia and P. inflata, but no wild
species carries the tandem.
RNAi as Antiviral Defense A System Plants Evolved Long Before We Discovered It
The CMV experiment in the Koseki study hints at something bigger: RNAi didn't evolve as a gene-
regulation tool. In plants, it is first and foremost an antiviral immune system.
Most plant viruses have RNA genomes that pass through a dsRNA intermediate during replication.
The plant cell recognizes this foreign dsRNA, processes it into siRNAs via Dicer, loads those
siRNAs into AGO proteins, and uses them to seek and destroy any RNA with matching sequence.
It's an adaptive, sequence-specific defense conceptually similar to CRISPR-Cas in bacteria, but
operating entirely at the RNA level.
Viruses fought back. Many plant viruses evolved viral suppressors of RNA silencing (VSRs)
proteins that interfere with specific steps of the pathway. The CMV 2b protein is one of the best
characterized. It operates through at least two distinct mechanisms: first, it binds directly to small
RNA duplexes (siRNA/siRNA* duplexes) along the duplex face, preventing their loading into AGO
proteins essentially intercepting the guide molecules before they can be incorporated into
functional RISC complexes. Second, 2b physically interacts with AGO1 through its PAZ-interacting
domain, blocking the slicer activity of preassembled RISC even when siRNAs are already loaded.
Additionally, 2b is imported into the nucleus, where it has been shown to interfere with the
production and systemic transport of the mobile silencing signal a 21–24 nt siRNA population that
normally moves through plasmodesmata and phloem to establish silencing in distant tissues ahead
of viral spread.
Other viral suppressors target different nodes of the pathway. The ** P19 protein of tombusviruses **
(such as Tomato bushy stunt virus) acts as a molecular caliper, binding siRNA duplexes in a size-
selective manner specifically sequestering 21-nt duplexes through contacts with the phosphate
backbone, effectively titrating out the Dicer products before they reach AGO. ** The P38 protein **
(turnip crinkle virus) mimics the ** GW motif found in endogenous AGO-interacting proteins likeGW182 ** ,
competitively displacing these cofactors and disrupting RISC assembly. The HC-Pro
protease of potyviruses (such as Tobacco etch virus) interferes with siRNA methylation by the
HEN1 methyltransferase unmethylated siRNAs are rapidly uridylated at their 3' ends and degraded,
effectively draining the siRNA pool. And the ** P6 protein of CaMV ** (Cauliflower mosaic virus the
same virus whose promoter was used in the original Jorgensen construct) suppresses silencing by
forming large inclusion bodies that sequester dsRNA and prevent Dicer from accessing it.
This molecular arms race between plant RNAi and viral counter-defense is what originally revealed
the pathway's importance. The very fact that viruses invest heavily in suppressing RNAi with some
viruses encoding multiple suppressors targeting different steps is proof of how effective it is as a
defense mechanism.
In mammals, the antiviral role of RNAi has been largely supplanted by the interferon system PKR,
OAS/RNase L, RIG-I, and MDA5 detect dsRNA and trigger innate immune responses instead. But
the core RNAi machinery (Dicer, AGO, RISC) has been retained and repurposed primarily for
endogenous gene regulation via microRNAs. Whether mammalian RNAi still plays a direct
antiviral role remains debated, but the evolutionary origin of the pathway as a defense against
parasitic nucleic acids is well established.Understanding this antiviral origin matters for therapeutic design: it explains why mammalian cells are inherently equipped to process and use synthetic siRNAs,
even though they no longer rely on RNAi for viral defense. The machinery is there it just needed a new purpose.
The Arc of Discovery
1990 A phenomenon is discovered by accident. Homologous genes silence each other coordinately,
reversibly, post-transcriptionally. No mechanism known.
1998 The trigger is identified: double-stranded RNA, acting catalytically.
2005 The same mechanism operates in nature, in a non-transgenic plant. siRNAs detected. Viral
suppressor reversal provides causal proof.
2012 The genomic architecture is revealed: tandem gene copies generating dsRNA, controlled by a
still-unknown regulatory locus.
Each paper answered the question left open by its predecessor.
__Why This Matters for Medicine__
FDA-approved RNAi drugs patisiran for hereditary transthyretin amyloidosis, givosiran for acute
hepatic porphyria, inclisiran for hypercholesterolemia all exploit the same dsRNA-triggered
pathway first glimpsed in white petunias in 1990.
The GalNAc-siRNA conjugate platform enables hepatocyte-specific delivery by exploiting
asialoglycoprotein receptor biology. Extrahepatic delivery to CNS, muscle, lung remains the
frontier, with lipid nanoparticles, antibody-siRNA conjugates, and exosome-based approaches under
active development.
The biology that a flower breeder stumbled upon 35 years ago is now a multi-billion-dollar
therapeutic modality. The distance from a puzzling white petunia to a patient receiving an RNA
drug is shorter than it looks.
